PAMAdam
PAMAdam (Parameter-wise Absolute Mean ADAM) is my new optimizer. It is very similar to LAMB, but it uses different function called absme:
def absme(t):
return t.abs().mean()
This absme will be applied on the parameter tensor and on the Adam step.
Let’s explain what absme will do in a diagram:
t = torch.randn(22)
q = t.abs()
m = q.mean()
def absme(t):
return t.abs().mean()
plt.scatter(range(0,22), t) #blue
plt.scatter(range(0,22), q+0.1) #orange
plt.scatter(range(0,1), m) #green
a = absme(t)
plt.scatter(range(2,3), a) #red
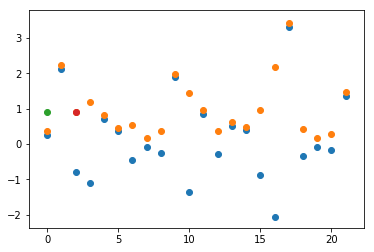
Blue dots are data from the normal distribution, orange dots are abs() of blues, green and red is the same absme value.
The function absme is to express the abs, mean operation.
Inside PAMAdam at the very end we will calculate absme two times:
absme1 = absme(p.data)
absme2 = absme(step)
Where p.data is the parameter, and the step is the Adam step. Note we removed the weight decay to simplify things.
Then the final update of param p would be:
p.data = p.data - lr*absme1/absme2 * step
Here is the full code:
class PAMAdam(Optimizer):
def __init__(self, params, lr=1e-3, betas=(0.9, 0.999), eps=1e-8):
defaults = dict(lr=lr, betas=betas, eps=eps)
super(PAMAdam, self).__init__(params, defaults)
def __setstate__(self, state):
super(PAMAdam, self).__setstate__(state)
def absme(self,t):
return t.abs().mean()
def step(self, closure=None):
for group in self.param_groups:
for p in group['params']:
if p.grad is None:
continue
grad = p.grad.data
state = self.state[p]
if len(state) == 0:
state['step'] = 0
state['agrad'] = torch.zeros_like(p.data) # grad average
state['agrad2'] = torch.zeros_like(p.data) # Hadamar grad average
state['step'] += 1
agrad, agrad2 = state['agrad'], state['agrad2']
beta1, beta2 = group['betas']
agrad.mul_(beta1).add_(1 - beta1, grad)
agrad2.mul_(beta2).addcmul_(1 - beta2, grad, grad)
bias_1 = 1 - beta1 ** state['step']
bias_2 = 1 - beta2 ** state['step']
agrad = agrad.div(bias_1)
agrad2 = agrad2.div(bias_2)
step = agrad / agrad2.sqrt().add_(group['eps'])
absme1 = self.absme(p.data)
absme2 = self.absme(step)
p.data.add_(-group['lr']*absme1/absme2, step)
return loss